Activación de STAT3 por hipoxia en modelos in vitro de cáncer de cuello uterino y en células endoteliales
Resumen
Introducción. El microambiente tumoral influye en el comportamiento de las células cancerosas. Especialmente, el estímulo de agentes estresantes, como la hipoxia, se convierte en un factor crítico para la evolución y el tratamiento del cáncer. La reacción celular frente a diversos estímulos se manifiesta en la activación de vías de señalización como la JAK/STAT, una de las más importantes por sus efectos en la diferenciación y proliferación celular.
Objetivo. Evaluar el estado de la vía JAK/STAT mediante la expresión o activación de la proteína STAT3 en células de cáncer de cuello uterino (HeLa) y en células endoteliales (EA.hy926) sometidas a hipoxia.
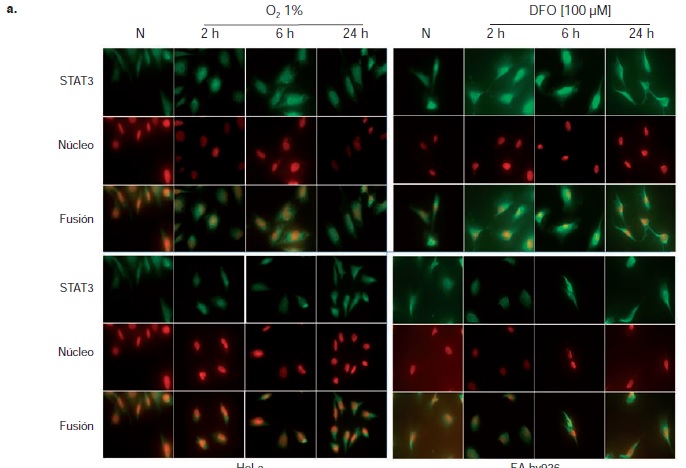
Materiales y métodos. Las líneas celulares se sometieron a condiciones de hipoxia física (1 % de O2) o química (100 μM de deferoxamina, DFO) durante dos, seis y 24 horas. Mediante Western blot se determinó el cambio en la expresión y activación de STAT3, y mediante inmunofluorescencia indirecta, su localización subcelular.
Resultados. La hipoxia se evidenció por la activación y translocación al núcleo del HIF-1. Ni la hipoxia física ni la química alteraron la expresión de STAT3, pero sí la activación, según se comprobó por su fosforilación y su translocación al núcleo en los dos modelos bajo estudio.
Conclusiones. Se evidenció la importancia de la hipoxia como un estímulo que modifica la activación de la proteína STAT3 en las células HeLa y EA.hy926, lo cual la convierte en un elemento importante en el diseño de estrategias terapéuticas contra el cáncer.
Descargas
Referencias bibliográficas
Varmus H. The new era in cancer research. Science. 2006;312:1162-5. http://dx.doi.org/10.1126/science.1126758
Vogelstein B, Kinzler KW. Cancer genes and the pathways they control. Nat Med. 2004;10:789-99. http://dx.doi.org/10.1038/nm1087
Caffarel MM, Coleman N. Oncostatin M receptor is a novel therapeutic target in cervical squamous cell carcinoma. J Pathol. 2014;232:386-90. http://dx.doi.org/10.1002/path.4305
Cao Y, DePinho RA, Ernst M, Vousden K. Cancer research: Past, present and future. Nat Rev Cancer. 2011;11:749-54. http://dx.doi.org/10.1038/nrc3138
Siemann DW. Tumor microenvironment. Chichester, UK.: John Wiley & Sons, Ltd; 2010. http://dx.doi.org/10.1002/9780470669891
Sullivan R, Graham CH. Hypoxia-driven selection of the metastatic phenotype. Cancer Metastasis Rev. 2007;26:319-31. http://dx.doi.org/10.1007/s10555-007-9062-2
Brahimi-Horn MC, Chiche J, Pouysségur J. Hypoxia and cancer. J Mol Med. 2007;85:1301-7. http://dx.doi.org/10.1007/s00109-007-0281-3
Liao D, Johnson RS. Hypoxia: A key regulator of angiogenesis in cancer. Cancer Metastasis Rev. 2007;26:281-90. http://dx.doi.org/10.1007/s10555-007-9066-y
Kenneth NS, Rocha S. Regulation of gene expression by hypoxia. Biochem J. 2008;414:19-29. http://dx.doi.org/10.1042/BJ20081055
Gadducci A, Guerrieri ME, Greco C. Tissue biomarkers as prognostic variables of cervical cancer. Crit Rev Oncol Hematol. 2013;86:104-29. http://dx.doi.org/10.1016/j.critrevonc. 2012.09.003
Schindler C, Levy DE, Decker T. JAK-STAT signaling: From interferons to cytokines. J Biol Chem. 2007;282:20059-63.http://dx.doi.org/10.1074/jbc.R700016200
Paukku K, Silvennoinen O. STATs as critical mediators of signal transduction and transcription: Lessons learned from STAT5. Cytokine Growth Factor Rev. 2004;15:435-55. http://dx.doi.org/10.1016/j.cytogfr.2004.09.001
Bowman T, García R, Turkson J, Jove R. STATs in oncogenesis. Oncogene. 2000;19:2474-88. http://dx.doi.org/10.1038/sj.onc.1203527
Takemoto S, Ushijima K, Kawano K, Yamaguchi T, Terada A, Fujiyoshi N, et al. Expression of activated signal transducer and activator of transcription-3 predicts poor prognosis in cervical squamous-cell carcinoma. Br J Cancer. 2009;101:967-72. http://dx.doi.org/10.1038/sj.bjc.6605212
Fan Z, Cui H, Xu X, Lin Z, Zhang X, Kang L, et al. MiR-125a suppresses tumor growth, invasion and metastasis in cervical cancer by targeting STAT3. Oncotarget. 2015;6:25266-80.http://dx.doi.org/10.18632/oncotarget.4457
Shukla S, Shishodia G, Mahata S, Hedau S, Pandey A, Bhambhani S, et al. Aberrant expression and constitutive activation of STAT3 in cervical carcinogenesis: Implications in high-risk human papillomavirus infection. Mol Cancer. 2010;9:282. http://dx.doi.org/10.1186/1476-4598-9-282
Grote K, Luchtefeld M, Schieffer B. JANUS under stress--role of JAK/STAT signaling pathway in vascular diseases. Vascul Pharmacol. 2005;43:357-63. http://dx.doi.org/10.1016/j.vph.2005.08.021
Dudley AC, Thomas D, Best J, Jenkins A. A VEGF/JAK2/STAT5 axis may partially mediate endothelial cell tolerance to hypoxia. Biochem J. 2005;390:427-36. http://dx.doi.org/10.1042/BJ20050351
Levy DE, Darnell JE. Stats: Transcriptional control and biological impact. Nat Rev Mol Cell Biol. 2002;3:651-62. http://dx.doi.org/10.1038/nrm909
Constantinescu SN, Girardot M, Pecquet C. Mining for JAK-STAT mutations in cancer. Trends Biochem Sci. 2008;33:122-31. http://dx.doi.org/10.1016/j.tibs.2007.12.002
Pensa S, Regis G, Boselli D, Novelli F, Poli V. STAT1 and STAT3 in tumorigenesis: Two sides of the same coin? En: Madame Curie Bioscience Database. Austin (TX): Landes Bioscience; 2000-2013.
Pilati C, Amessou M, Bihl MP, Balabaud C, Nhieu JT, Paradis V, et al. Somatic mutations activating STAT3 in human inflammatory hepatocellular adenomas. J Exp Med. 2011;208:1359-66. http://dx.doi.org/10.1084/jem.20110283
Yu H, Kortylewski M, Pardoll D. Crosstalk between cancer and immune cells: Role of STAT3 in the tumour microenvironment. Nat Rev Immunol. 2007;7:41-51. http://dx.doi.org/10.1038/nri1995
Grivennikov SI, Karin M. Dangerous liaisons: STAT3 and NF-kappaB collaboration and crosstalk in cancer. Cytokine Growth Factor Rev. 2010;21:11-9. http://dx.doi.org/10.1016/j.cytogfr.2009.11.005
Mauge L, Terme M, Tartour E, Helley D. Control of the adaptive immune response by tumor vasculature. Front Oncol. 2014;4:61. http://dx.doi.org/10.3389/fonc.2014.00061
Pawlus MR, Wang L, Hu C-J. STAT3 and HIF1alpha cooperatively activate HIF1 target genes in MDA-MB-231 and RCC4 cells. Oncogene. 2014;33:1670-9. http://dx.doi.org/10.1038/onc.2013.115
Demaria M, Giorgi C, Lebiedzinska M, Esposito G, D’angeli L, Bartoli A, et al. A STAT3-mediated metabolic switch is involved in tumour transformation and STAT3 addiction. Aging (Albany NY). 2010;2:823-42. http://dx.doi.org/10.18632/aging.100232
Wei L-H, Kuo M-L, Chen C-A, Chou C-H, Lai K-B, Lee C-N, et al. Interleukin-6 promotes cervical tumor growth by VEGF-dependent angiogenesis via a STAT3 pathway. Oncogene. 2003;22:1517-27. http://dx.doi.org/10.1038/sj.onc.1206226
Martorell L, Gentile M, Rius J, Rodríguez C, Crespo J, Badimon L, et al. The hypoxia-inducible factor 1/NOR-1 axis regulates the survival response of endothelial cells to hypoxia. Mol Cell Biol. 2009;29:5828-42. http://dx.doi.org/10.1128/MCB.00945-09
Guo M, Song L-P, Jiang Y, Liu W, Yu Y, Chen G-Q. Hypoxia-mimetic agents desferrioxamine and cobalt chloride induce leukemic cell apoptosis through different hypoxiainducible factor-1alpha independent mechanisms. Apoptosis. 2006;11:67-77. http://dx.doi.org/10.1007/s10495-005-3085-3
Joyce JA. Therapeutic targeting of the tumor microenvironment. Cancer Cell. 2005;7:513-20. http://dx.doi.org/10.1016/j.ccr.2005.05.024
Kaelin WGJ, Ratcliffe PJ. Oxygen sensing by metazoans: The central role of the HIF hydroxylase pathway. Mol Cell. 2008;30:393-402. http://dx.doi.org/10.1016/j.molcel.2008.04.009
Zhu P, Ning Y, Yao L, Chen M, Xu C. The proliferation, apoptosis, invasion of endothelial-like epithelial ovarian cancer cells induced by hypoxia. J Exp Clin Cancer Res. 2010;29:124. http://dx.doi.org/10.1186/1756-9966-29-124
Liu L, McBride KM, Reich NC. STAT3 nuclear import is independent of tyrosine phosphorylation and mediated by importin-alpha3. Proc Natl Acad Sci USA. 2005;102:8150-5.http://dx.doi.org/10.1073/pnas.0501643102
Levy DE. The house that JAK/STAT built. Cytokine Growth Factor Rev.1997;8:81-90. http://dx.doi.org/10.1016/S1359-6101(96)00054-8
Okamoto W, Okamoto I, Arao T, Yanagihara K, Nishio K, Nakagawa K. Differential roles of STAT3 depending on the mechanism of STAT3 activation in gastric cancer cells. Br J Cancer. 2011;105:407-12. http://dx.doi.org/10.1038/bjc.2011.246
Bauer J, Margolis M, Schreiner C, Edgell CJ, Azizkhan J, Lazarowski E, et al. In vitro model of angiogenesis using a human endothelium-derived permanent cell line: Contributions of induced gene expression, G-proteins, and integrins. J Cell Physiol. 1992;153:437-49. http://dx.doi.org/10.1002/jcp.1041530302
Schutz A, Roser K, Klitzsch J, Lieder F, Aberger F, Gruber W, et al. Lung adenocarcinomas and lung cancer cell lines show association of MMP-1 expression with STAT3 activation. Transl Oncol. 2015;8:97-105. http://dx.doi.org/10.1016/j.tranon.2015.02.002
Bertout JA, Patel SA, Simon MC. The impact of O2 availability on human cancer. Nat Rev Cancer. 2008;8:967-75. http://dx.doi.org/10.1038/nrc2540
Li WX. Canonical and non-canonical JAK-STAT signaling. Trends Cell Biol. 2008;18:545-51. http://dx.doi.org/10.1016/j.tcb.2008.08.008
Wen Z, Zhong Z, Darnell JE. Maximal activation of transcription by Stat1 and Stat3 requires both tyrosine and serine phosphorylation. Cell. 1995;82:241-50.
Yuan Z-L, Guan Y-J, Chatterjee D, Chin YE. STAT3 dimerization regulated by reversible acetylation of a single lysine residue. Science. 2005;307:269-73. http://dx.doi.org/10.1126/science.1105166
Jung JE, Lee HG, Cho IH, Chung DH, Yoon S-H, Yang YM, et al. STAT3 is a potential modulator of HIF-1-mediated VEGF expression in human renal carcinoma cells. FASEB J. 2005;19:1296-8. http://dx.doi.org/10.1096/fj.04-3099fje
Xu Q, Briggs J, Park S, Niu G, Kortylewski M, Zhang S, et al. Targeting Stat3 blocks both HIF-1 and VEGF expression induced by multiple oncogenic growth signaling pathways. Oncogene. 2005;24:5552-60. http://dx.doi.org/10.1038/sj.onc.1208719
Darnell JE. STAT3, HIF-1, glucose addiction and Warburg effect. Aging (Albany NY). 2010;2:890-1. http://dx.doi.org/10.18632/aging.100239
Jung JE, Kim HS, Lee CS, Shin YJ, Kim YN, Kang GH, et al. STAT3 inhibits the degradation of HIF-1alpha by pVHLmediated ubiquitination. Exp Mol Med. 2008;40:479-85.
http://dx.doi.org/10.3858/emm.2008.40.5.479
Hirsila M, Koivunen P, Xu L, Seeley T, Kivirikko KI, Myllyharju J. Effect of desferrioxamine and metals on the hydroxylases in the oxygen sensing pathway. FASEB J. 2005;19:1308-10. http://dx.doi.org/10.1096/fj.04-3399fje
Chen H, Guan Y, Yuan G, Zhang Q, Jing N. A perylene derivative regulates HIF-1alpha and Stat3 signaling pathways. Bioorg Med Chem. 2014;22:1496-505. http://dx.doi.org/10.1016/j.bmc.2013.10.018
Demaria M, Poli V. From the nucleus to the mitochondria and back: The odyssey of a multitask STAT3. Cell Cycle. 2011;10:3221-2. http://dx.doi.org/10.4161/cc.10.19.17379
Blaskovich MA, Sun J, Cantor A, Turkson J, Jove R, Sebti SM. Discovery of JSI-124 (cucurbitacin I), a selective Janus kinase/signal transducer and activator of transcription 3 signaling pathway inhibitor with potent antitumor activity against human and murine cancer cells in mice. Cancer Res. 2003;63:1270-9.
Yu MO, Park K-J, Park D-H, Chung Y-G, Chi S-G, Kang S-H. Reactive oxygen species production has a critical role in hypoxia-induced Stat3 activation and angiogenesis in human glioblastoma. J Neurooncol. 2015;125:55-63. http://dx.doi.org/10.1007/s11060-015-1889-8
Algunos artículos similares:
- Constanza Pardo, Ricardo Cendales, Supervivencia de pacientes con cáncer de cuello uterino tratadas en el Instituto Nacional de Cancerología , Biomédica: Vol. 29 Núm. 3 (2009)
- Raúl Murillo, Ricardo Cendales, Carolina Wiesner, Marion Piñeros, Sandra Tovar, Efectividad de la citología cérvico-uterina para la detección temprana de cáncer de cuello uterino en el marco del sistema de salud de Colombia , Biomédica: Vol. 29 Núm. 3 (2009)
- Ricardo Cendales, Carolina Wiesner, Raúl Hernando Murillo, Marion Piñeros, Sandra Tovar, Juan Carlos Mejía, La calidad de las citologías para tamización de cáncer de cuello uterino en cuatro departamentos de Colombia: un estudio de concordancia , Biomédica: Vol. 30 Núm. 1 (2010)
- Miguel Amézquita , Geny Carolina Silva , Diego Antonio Restrepo , Linda Margarita Ibata , Rafael Niño , Maximiliano Bustacara , Víctor Alexander Sáenz , Dieric Anderson Díaz , Milena Alarcón , Luz Adriana Quintero , Análisis del impacto presupuestal en Colombia de la prueba de HPV con genotipificación comparada con la citología , Biomédica: Vol. 42 Núm. 2 (2022)
| Estadísticas de artículo | |
|---|---|
| Vistas de resúmenes | |
| Vistas de PDF | |
| Descargas de PDF | |
| Vistas de HTML | |
| Otras vistas | |